Forward-looking: Companies and research institutions have been exploring DNA as a potential "storage system" for archiving digital data for a long time. A new approach is now emerging, offering additional financial incentives by eliminating the need for complex and expensive lab-created DNA strands from scratch. This novel variation in the field shows promise in terms of cost-effectiveness and simplicity.
Universally known as DNA, the deoxyribonucleic acid polymer is the fundamental building block that governs every living organism we have discovered to date. It serves as the control center for cellular activity and determines the biological traits expressed by animals, plants, and all other life forms on our planet. Genetic instructions within an organism's DNA are encoded using a four-base system, consisting of adenine (A), thymine (T), guanine (G), and cytosine (C) nucleotides.
This simple yet universally valid approach to genetic encoding has long fascinated scientists and companies worldwide. The biological model could provide an answer to challenges in storing vast amounts of digital data generated by the internet and the numerous devices that populate our all-digital global society.
Experiments involving the storage of digital bits in DNA strands are typically expensive due to the need for creating new synthetic DNA sequences through complex and error-prone processes. However, a team of researchers, led by Cheng Kai Lim, a synthetic biologist at the National University of Singapore, is now proposing an alternative approach. This innovative method aims to address the cost issue by utilizing "living" DNA strands.
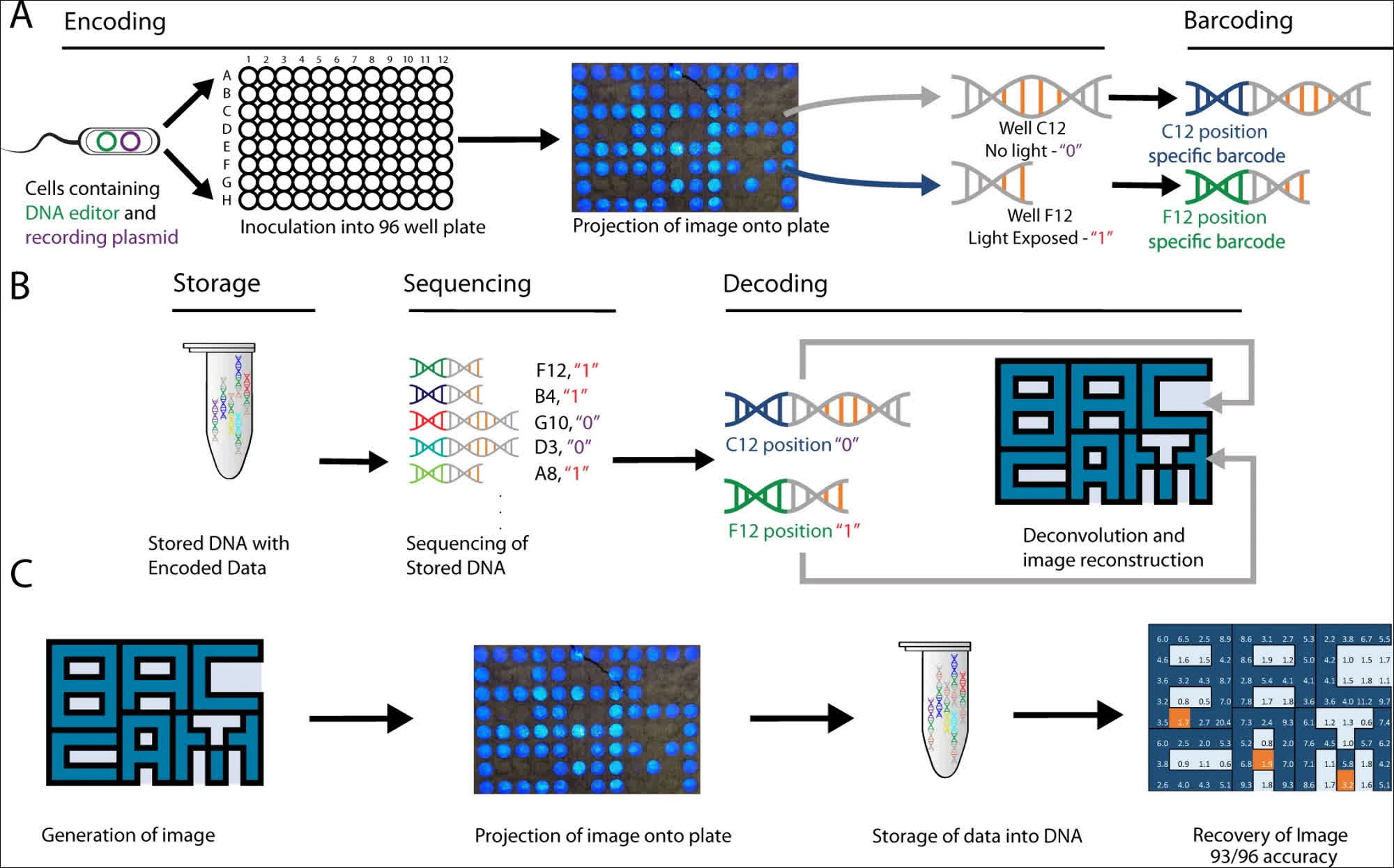
The researchers utilized the DNA capabilities present in the cells of Escherichia coli, a coliform bacterium commonly found in the digestive tracts of warm-blooded organisms, including humans. These DNA strands are equipped with "optogenetic" circuits capable of detecting the presence or absence of light. Leveraging this functionality, the researchers developed BacCam, a biological camera that can directly capture and store images within DNA.
The team devised a method for capturing "2-dimensional light patterns" in DNA by leveraging optogenetic circuits to record light exposure and encode spatial locations using barcoding. The researchers explained that the stored images were subsequently retrieved through "high-throughput next-generation sequencing," which opens the door to "integrating biological systems with digital devices."
The researchers were able to store simple 96-bit images for a total of 1152 bits in biological DNA, successfully demonstrating the "multiplexing" ability of their method by capturing and storing two different images at the same time, using red and blue light sources. The DNA-stored images could be recovered with an accuracy of at least 90 percent for all images, which is a significant achievement in biology-based digital archives. However, it falls short of the reliability required for a robust file storage system.
Nonetheless, the study suggests that a DNA pool can store and retrieve between 100 and 1,000 different images in a single run. The researchers are optimistic about the ongoing advancements in DNA data storage, as their method showcases additional applications of this promising technology.